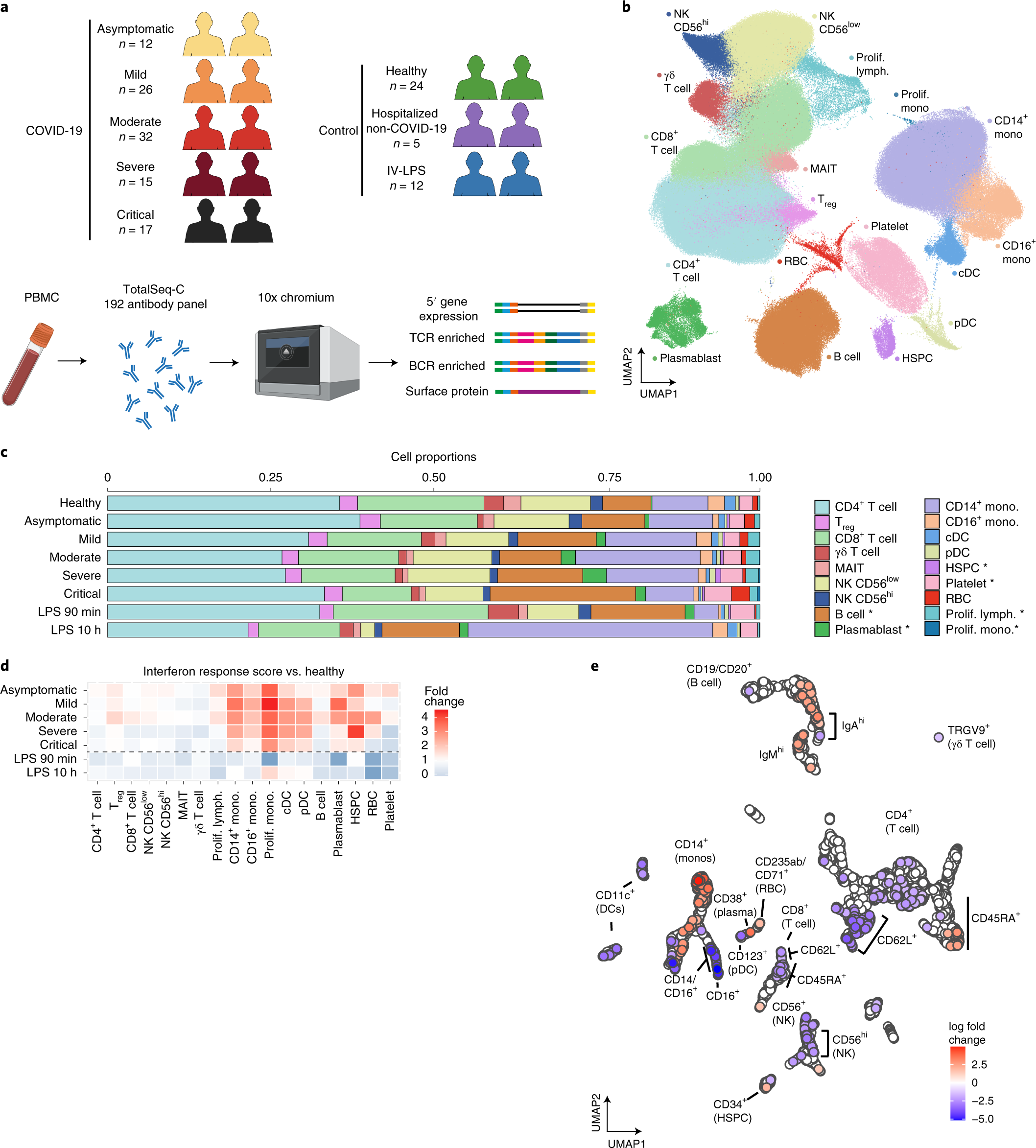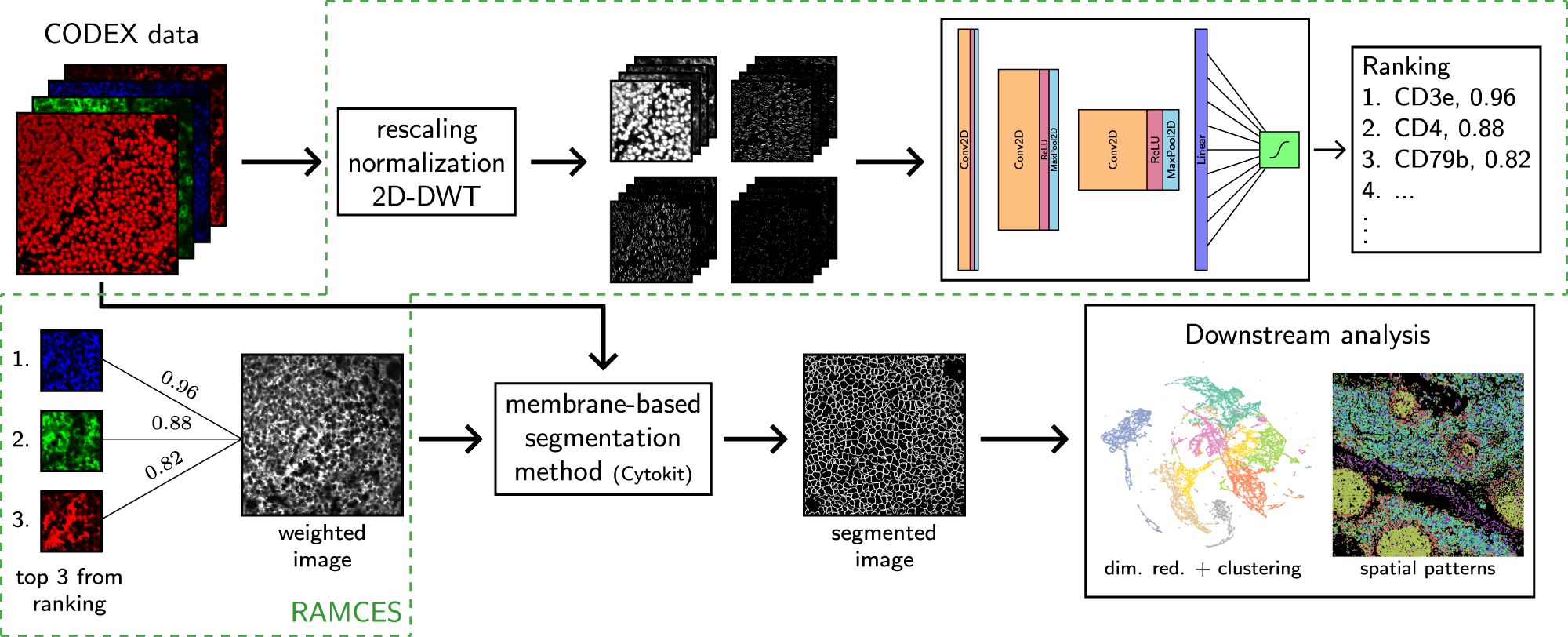
Membrane marker selection for segmenting single cell spatial proteomics data | Nature Communications
CCS/MSP430FR2033: EXPECTED AN IDENTIFIER ERROR IN CCS - Code Composer Studio forum - Code Composer Studio™︎ - TI E2E support forums

syntax error, unexpected end of file, expecting function (T_FUNCTION) or const (T_CONST) · Issue #5434 · craftcms/cms · GitHub

A multi-marker association method for genome-wide association studies without the need for population structure correction | Nature Communications

pset1 - Mario error: expected ';' at end of declaration...all solutions i've found fail - CS50 Stack Exchange
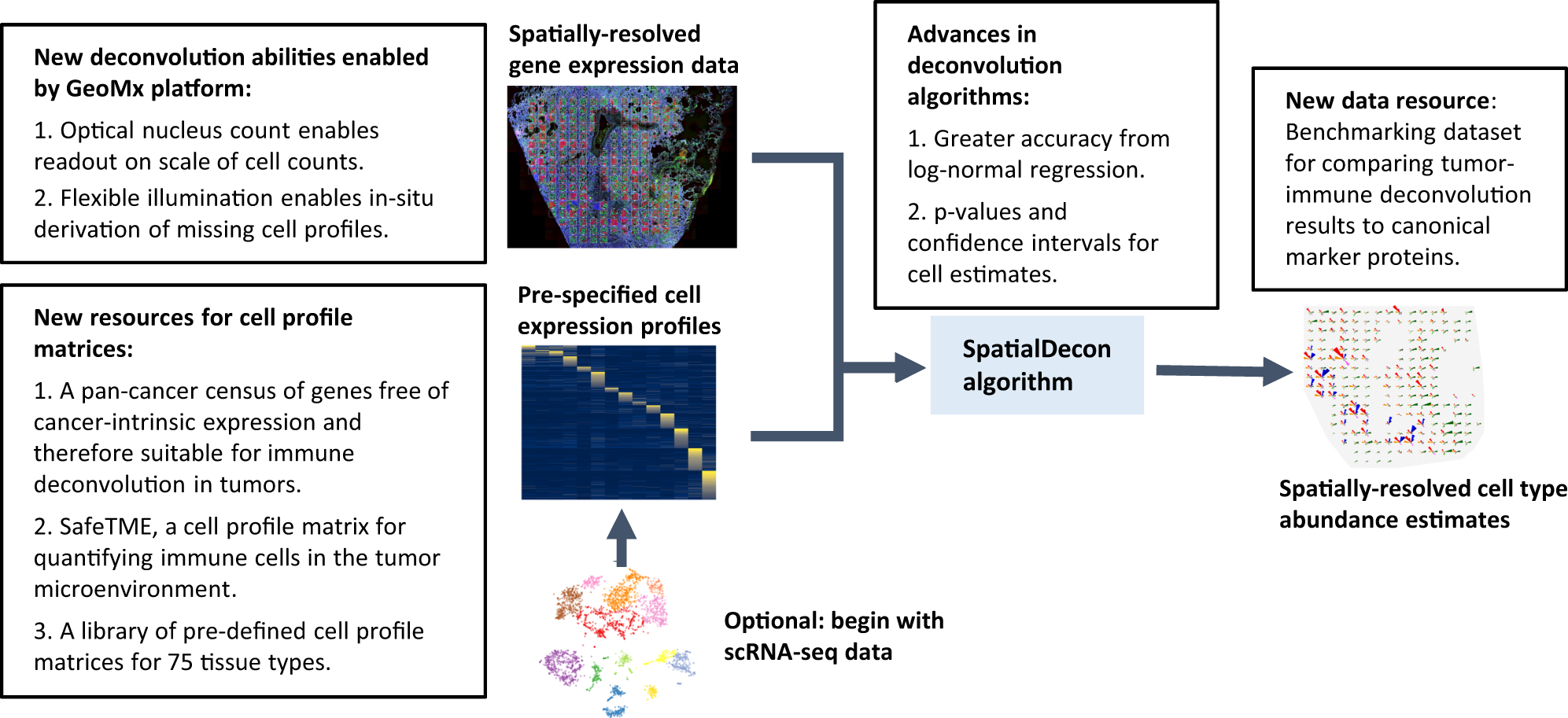
Advances in mixed cell deconvolution enable quantification of cell types in spatial transcriptomic data | Nature Communications

pset1 - Mario error: expected ';' at end of declaration...all solutions i've found fail - CS50 Stack Exchange
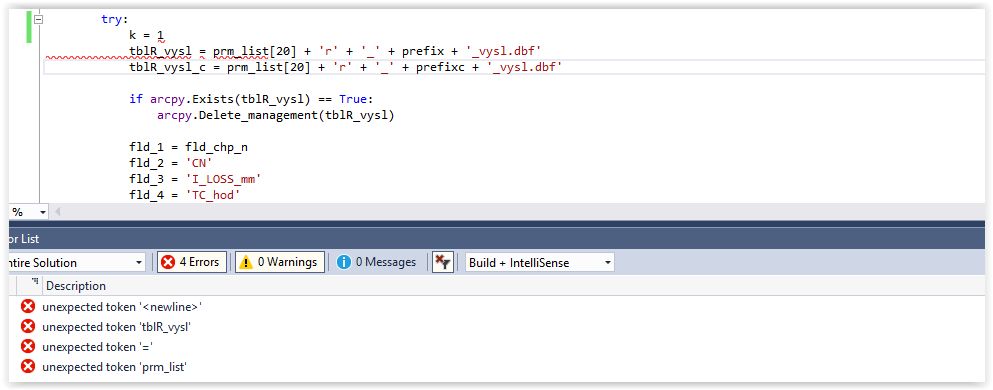
Editing code leads to unexpected token '<newline>' errors in error list · Issue #4835 · microsoft/PTVS · GitHub

Age-related brain atrophy is not a homogenous process: Different functional brain networks associate differentially with aging and blood factors | PNAS

Multi-marker DNA metabarcoding detects suites of environmental gradients from an urban harbour | Scientific Reports

Settings has an invalid type, expected "string". Fix in JSON. Live Server Not Previewing. · Issue #97355 · microsoft/vscode · GitHub

Estimated accuracy of genomic prediction (SE) at different levels of... | Download Scientific Diagram

c++ - GDI+ library causes "error C2760: syntax error: unexpected token 'identifier', expected 'type specifier'" in VS2017 when compiled for XP - Stack Overflow


![Solved //Error message: "[Error] expected declaration | Chegg.com Solved //Error message: "[Error] expected declaration | Chegg.com](https://d2vlcm61l7u1fs.cloudfront.net/media%2F1bd%2F1bd5ba93-dee4-4c7d-8170-8bbd91da4eda%2FphpuE52EV.png)